DNA Sequencing Explained: Technologies and Applications
What is DNA Sequencing?
DNA sequencing is the process of determining the precise order of nucleotides within a DNA molecule. It involves reading the sequence of the four DNA bases - adenine (A), guanine (G), cytosine (C), and thymine (T) - that make up an organism's genetic code. This genetic information is essential for understanding the function, structure, and evolution of organisms, as well as for various applications in fields such as medicine, agriculture, and forensics.
The Evolution of DNA Sequencing Technologies
DNA sequencing technologies have undergone significant advancements over the years, enabling faster, more accurate, and cost-effective sequencing of genomes:
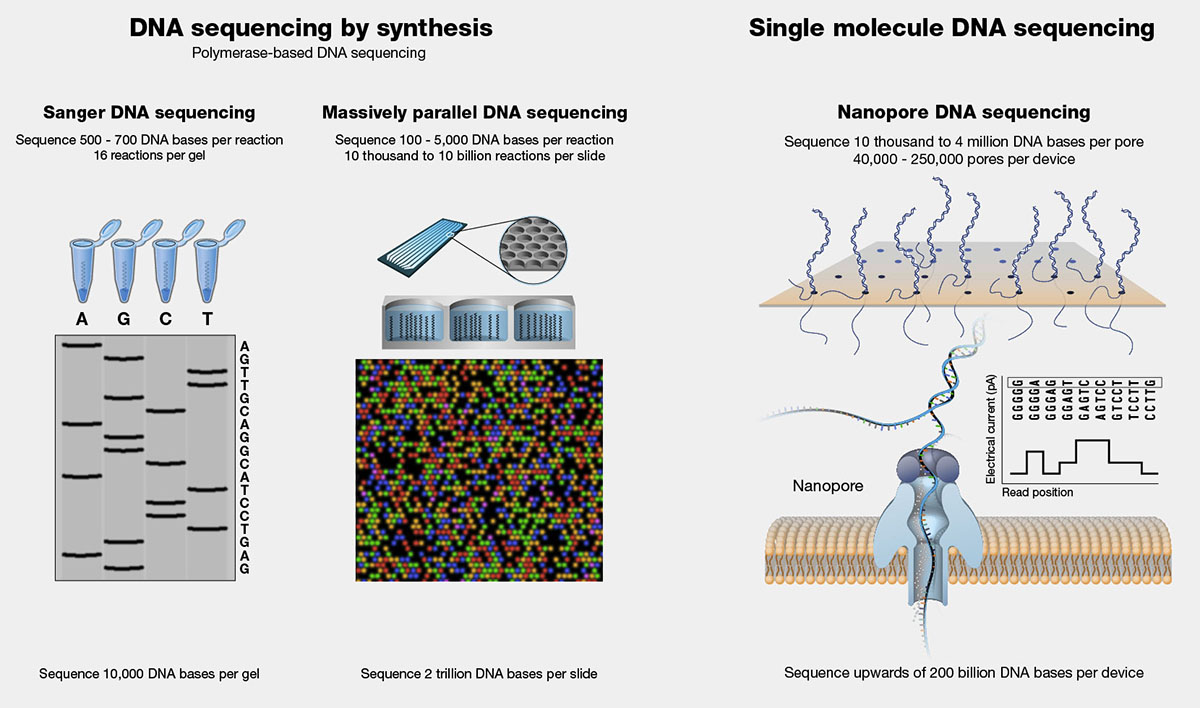
First-Generation Sequencing (Sanger Sequencing)
Developed by Frederick Sanger in the 1970s, Sanger sequencing was the first widely used method for DNA sequencing. This method involves the synthesis of DNA strands complementary to the template DNA, using modified nucleotides that terminate the synthesis at specific bases. The resulting DNA fragments are then separated by size using gel electrophoresis, and the sequence is determined by analyzing the pattern of bands on the gel.
Next-Generation Sequencing (NGS)
Next-generation sequencing technologies, introduced in the mid-2000s, revolutionized the field of genomics by enabling high-throughput, parallelized sequencing of DNA. NGS methods typically involve the fragmentation of DNA into short segments, followed by the simultaneous sequencing of millions of these fragments. Some common NGS platforms include Illumina (sequencing by synthesis), Ion Torrent (semiconductor sequencing), and Pacific Biosciences (single-molecule real-time sequencing).
Third-Generation Sequencing (Long-Read Sequencing)
Third-generation sequencing technologies, such as those developed by Oxford Nanopore Technologies and Pacific Biosciences, allow for the sequencing of long, continuous DNA fragments in real-time. These methods overcome some of the limitations of short-read sequencing, such as the difficulty in assembling complex genomes and resolving repetitive regions. Long-read sequencing enables the identification of structural variations, epigenetic modifications, and full-length transcript isoforms.
Applications of DNA Sequencing
DNA sequencing has a wide range of applications across various fields:
Genomics and Molecular Biology
DNA sequencing is the foundation of genomics research, enabling the study of complete genomes, gene function, and evolutionary relationships between organisms. It allows for the identification of genetic variations, mutations, and regulatory elements that contribute to phenotypic diversity and disease susceptibility.
Medical Diagnostics and Personalized Medicine
DNA sequencing plays a crucial role in the diagnosis and treatment of genetic disorders, cancer, and infectious diseases. By sequencing patient genomes or specific disease-associated genes, clinicians can identify causative mutations, predict disease risk, and guide personalized treatment strategies. Pharmacogenomics, which involves tailoring drug therapies based on an individual's genetic profile, relies heavily on DNA sequencing data.
Agriculture and Biotechnology
DNA sequencing is used to study the genomes of crops and livestock, enabling the identification of genes associated with desirable traits such as yield, disease resistance, and nutritional quality. This information can be used to develop improved varieties through marker-assisted selection or genetic engineering. Sequencing also facilitates the discovery of novel enzymes, biofuels, and biomaterials from microbial genomes.
Forensics and Ancestry
DNA sequencing is a powerful tool in forensic investigations, allowing for the identification of individuals based on trace amounts of genetic material. By comparing DNA profiles from crime scene samples with reference databases, investigators can link suspects to crimes or exonerate the innocent. DNA sequencing is also used in ancestry testing, where an individual's DNA is analyzed to determine their genetic lineage and geographical origins.
Challenges and Future Perspectives
Despite the remarkable advancements in DNA sequencing technologies, several challenges remain. The generation and analysis of massive amounts of sequencing data require robust computational infrastructure and bioinformatics tools. Interpreting the functional significance of genetic variations and integrating multi-omics data to gain a comprehensive understanding of biological systems are ongoing challenges.
Future developments in DNA sequencing aim to further improve accuracy, speed, and cost-effectiveness. Emerging technologies, such as nanopore sequencing and single-molecule sequencing, hold promise for real-time, portable, and ultra-long-read sequencing. The integration of DNA sequencing with other technologies, such as genome editing, single-cell analysis, and synthetic biology, will open up new avenues for scientific discovery and biotechnological applications.
Further Reading
Human Genomics, The application of long-read sequencing in clinical settings
Computational and Structural Biotechnology Journal, Long walk to genomics: History and current approaches to genome sequencing and assembly